GROMACS 2020 was released on January 1, 2020. Patch releases may have been made since then, please use the updated versions! Here are some highlights, along with more detail in the links below!
There are several useful performance improvements, with or without GPUs, all enabled and automated by default. In addition, several new features are available for running simulations. The new features include:
- Density-guided simulations allow "fitting" atoms into three-dimensional density maps.
- Inclusion of gmxapi 0.1, an API and user interface for managing complex simulations, data flow, and pluggable molecular dynamics extension code.
- New modular simulator can be built from individual objects describing different calculations happening at each simulation step.
- Parrinello-Rahman pressure coupling is now also available for the md-vv integrator.
- Running almost the entire simulation step on a single CUDA compatible GPU for supported types of simulations, including coordinate update and constraint calculation.
Click here to view the full Release notes.
What is GROMACS?
GROMACS is a versatile package to perform molecular dynamics, i.e. simulate the Newtonian equations of motion for systems with hundreds to millions of particles.
It is primarily designed for biochemical molecules like proteins, lipids, and nucleic acids that have a lot of complicated bonded interactions, but since GROMACS is extremely fast at calculating the non-bonded interactions (that usually dominate simulations) many groups are also using it for research on non-biological systems, e.g. polymers.
Creating Faster Molecular Dynamics Simulations with GROMACS 2020
The long-term collaboration between NVIDIA and the core GROMACS developers has delivered a simulation package for biomolecular systems that performs incredibly fast. Previous GROMACS packages saw a spike in performance using GPU acceleration, but at a computational expense, especially when using multiple GPUs to run a single simulation. The new performance features available in GROMACS 2020 address these issues, and now for many typical simulations, the entire timestep can run on the GPU, avoiding CPU and PCIe bottlenecks. Inter-GPU communication operations can now operate directly between GPU memory spaces, resulting in large performance improvements.
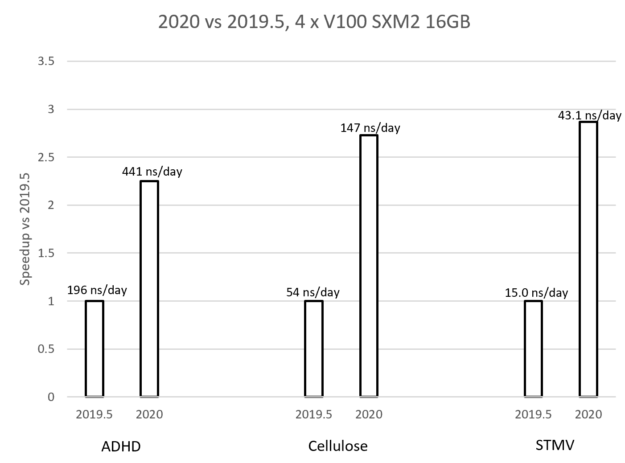
Figure 7: Comparison of GROMACS 2019 to GROMACS 2020 using three multi-GPU simulation examples.
You can read more details on the NVIDIA blog.
GROMACS 2020 Performance Improvements
- Up to a factor 2.5 speed-up of the non-bonded free-energy kernel - The non-bonded free-energy kernel is a factor 2.5 faster with non-zero A and B states and a factor 1.5 with one zero states. This especially improves the run performance when non-perturbed non-bonds are offloaded to a GPU. In that case, the PME-mesh calculation now always takes the most CPU time.
- Proper dihedrals of Fourier type and improper dihedrals of periodic type are SIMD accelerated
- Avoid configuring the own-FFTW with AVX512 enabled when GROMACS does not use AVX512 - Previously if GROMACS was configured to use any AVX flavor, the internally built FFTW would be configured to also contain AVX512 kernels. This could cause performance loss if the (often noisy) FFTW auto-tuner picks an AVX512 kernel in a run that otherwise only uses AVX/AVX2 which could run at higher CPU clocks without AVX512 clock speed limitation. Now AVX512 is only used for the internal FFTW if GROMACS is also configured with the same SIMD flavor.
- Update and constraints can run on a GPU - For standard simulations (see the user guide for more details), updates and constraints can be offloaded to a GPU with CUDA. Thus all compute-intensive parts of a simulation can be offloaded, which provides better performance when using a fast GPU combined with a slow CPU. By default, the update will run on the CPU, to use GPU in single rank simulations, one can use new '-update GPU' command-line option. For use with domain decomposition, please see below.
- GPU Direct Communications - When running on multiple GPUs with CUDA, communication operations can now be performed directly between GPU memory spaces (automatically routed, including via NVLink where available). This behavior is not yet enabled by default: the new codepaths have been verified by the standard GROMACS regression tests, but (at the time of release) still lack substantial "real-world" testing. They can be enabled by setting the following environment variables to any non-NULL value in your shell: GMX_GPU_DD_COMMS (for halo exchange communications between PP tasks); GMX_GPU_PME_PP_COMMS (for communications between PME and PP tasks); GMX_FORCE_UPDATE_DEFAULT_GPU can also be set in order to combine with the new GPU update feature (above). The combination of these will (for many common simulations) keep data resident on the GPU across most timesteps, avoiding expensive data transfers. Note that these currently require GROMACS to be built with its internal thread-MPI library rather than any external MPI library, and are limited to a single compute node. We stress that users should carefully verify results against the default path, and any reported issues will be gratefully received to help us mature the software.
- Bonded kernels on GPU have been fused - Instead of launching one GPU kernel for each listed interaction type, there is now one GPU kernel that handles all listed interactions. This improves performance when running bonded calculations on a GPU.
- The delay for ramp-up added to PP-PME tuning - Modern CPUs and GPUs can take a few seconds to ramp up their clock speeds. Therefore the PP-PME load balancing now starts after 5 seconds instead of after a few MD steps. This avoids sub-optimal performance settings.
Download the complete full documentation to learn more: http://manual.gromacs.org/2020/manual-2020.pdf
Let us know if you have any questions about getting a system optimized for GROMACS, or upgrading your current one.

GROMACS 2020 Highlights
GROMACS 2020 was released on January 1, 2020. Patch releases may have been made since then, please use the updated versions! Here are some highlights, along with more detail in the links below!
There are several useful performance improvements, with or without GPUs, all enabled and automated by default. In addition, several new features are available for running simulations. The new features include:
- Density-guided simulations allow "fitting" atoms into three-dimensional density maps.
- Inclusion of gmxapi 0.1, an API and user interface for managing complex simulations, data flow, and pluggable molecular dynamics extension code.
- New modular simulator can be built from individual objects describing different calculations happening at each simulation step.
- Parrinello-Rahman pressure coupling is now also available for the md-vv integrator.
- Running almost the entire simulation step on a single CUDA compatible GPU for supported types of simulations, including coordinate update and constraint calculation.
Click here to view the full Release notes.
What is GROMACS?
GROMACS is a versatile package to perform molecular dynamics, i.e. simulate the Newtonian equations of motion for systems with hundreds to millions of particles.
It is primarily designed for biochemical molecules like proteins, lipids, and nucleic acids that have a lot of complicated bonded interactions, but since GROMACS is extremely fast at calculating the non-bonded interactions (that usually dominate simulations) many groups are also using it for research on non-biological systems, e.g. polymers.
Creating Faster Molecular Dynamics Simulations with GROMACS 2020
The long-term collaboration between NVIDIA and the core GROMACS developers has delivered a simulation package for biomolecular systems that performs incredibly fast. Previous GROMACS packages saw a spike in performance using GPU acceleration, but at a computational expense, especially when using multiple GPUs to run a single simulation. The new performance features available in GROMACS 2020 address these issues, and now for many typical simulations, the entire timestep can run on the GPU, avoiding CPU and PCIe bottlenecks. Inter-GPU communication operations can now operate directly between GPU memory spaces, resulting in large performance improvements.
Figure 7: Comparison of GROMACS 2019 to GROMACS 2020 using three multi-GPU simulation examples.
You can read more details on the NVIDIA blog.
GROMACS 2020 Performance Improvements
- Up to a factor 2.5 speed-up of the non-bonded free-energy kernel - The non-bonded free-energy kernel is a factor 2.5 faster with non-zero A and B states and a factor 1.5 with one zero states. This especially improves the run performance when non-perturbed non-bonds are offloaded to a GPU. In that case, the PME-mesh calculation now always takes the most CPU time.
- Proper dihedrals of Fourier type and improper dihedrals of periodic type are SIMD accelerated
- Avoid configuring the own-FFTW with AVX512 enabled when GROMACS does not use AVX512 - Previously if GROMACS was configured to use any AVX flavor, the internally built FFTW would be configured to also contain AVX512 kernels. This could cause performance loss if the (often noisy) FFTW auto-tuner picks an AVX512 kernel in a run that otherwise only uses AVX/AVX2 which could run at higher CPU clocks without AVX512 clock speed limitation. Now AVX512 is only used for the internal FFTW if GROMACS is also configured with the same SIMD flavor.
- Update and constraints can run on a GPU - For standard simulations (see the user guide for more details), updates and constraints can be offloaded to a GPU with CUDA. Thus all compute-intensive parts of a simulation can be offloaded, which provides better performance when using a fast GPU combined with a slow CPU. By default, the update will run on the CPU, to use GPU in single rank simulations, one can use new '-update GPU' command-line option. For use with domain decomposition, please see below.
- GPU Direct Communications - When running on multiple GPUs with CUDA, communication operations can now be performed directly between GPU memory spaces (automatically routed, including via NVLink where available). This behavior is not yet enabled by default: the new codepaths have been verified by the standard GROMACS regression tests, but (at the time of release) still lack substantial "real-world" testing. They can be enabled by setting the following environment variables to any non-NULL value in your shell: GMX_GPU_DD_COMMS (for halo exchange communications between PP tasks); GMX_GPU_PME_PP_COMMS (for communications between PME and PP tasks); GMX_FORCE_UPDATE_DEFAULT_GPU can also be set in order to combine with the new GPU update feature (above). The combination of these will (for many common simulations) keep data resident on the GPU across most timesteps, avoiding expensive data transfers. Note that these currently require GROMACS to be built with its internal thread-MPI library rather than any external MPI library, and are limited to a single compute node. We stress that users should carefully verify results against the default path, and any reported issues will be gratefully received to help us mature the software.
- Bonded kernels on GPU have been fused - Instead of launching one GPU kernel for each listed interaction type, there is now one GPU kernel that handles all listed interactions. This improves performance when running bonded calculations on a GPU.
- The delay for ramp-up added to PP-PME tuning - Modern CPUs and GPUs can take a few seconds to ramp up their clock speeds. Therefore the PP-PME load balancing now starts after 5 seconds instead of after a few MD steps. This avoids sub-optimal performance settings.
Download the complete full documentation to learn more: http://manual.gromacs.org/2020/manual-2020.pdf
Let us know if you have any questions about getting a system optimized for GROMACS, or upgrading your current one.